CD8+ T cells utilize highly dynamic enhancer repertoires and regulatory
circuitry in response to infections
He B, Xing S, Chen C, Gao P, Teng L, Shan Q, Gullicksrud JA, Martin MD, Yu S, Harty JT, Badovinac VP, Tan K, and Xue HH
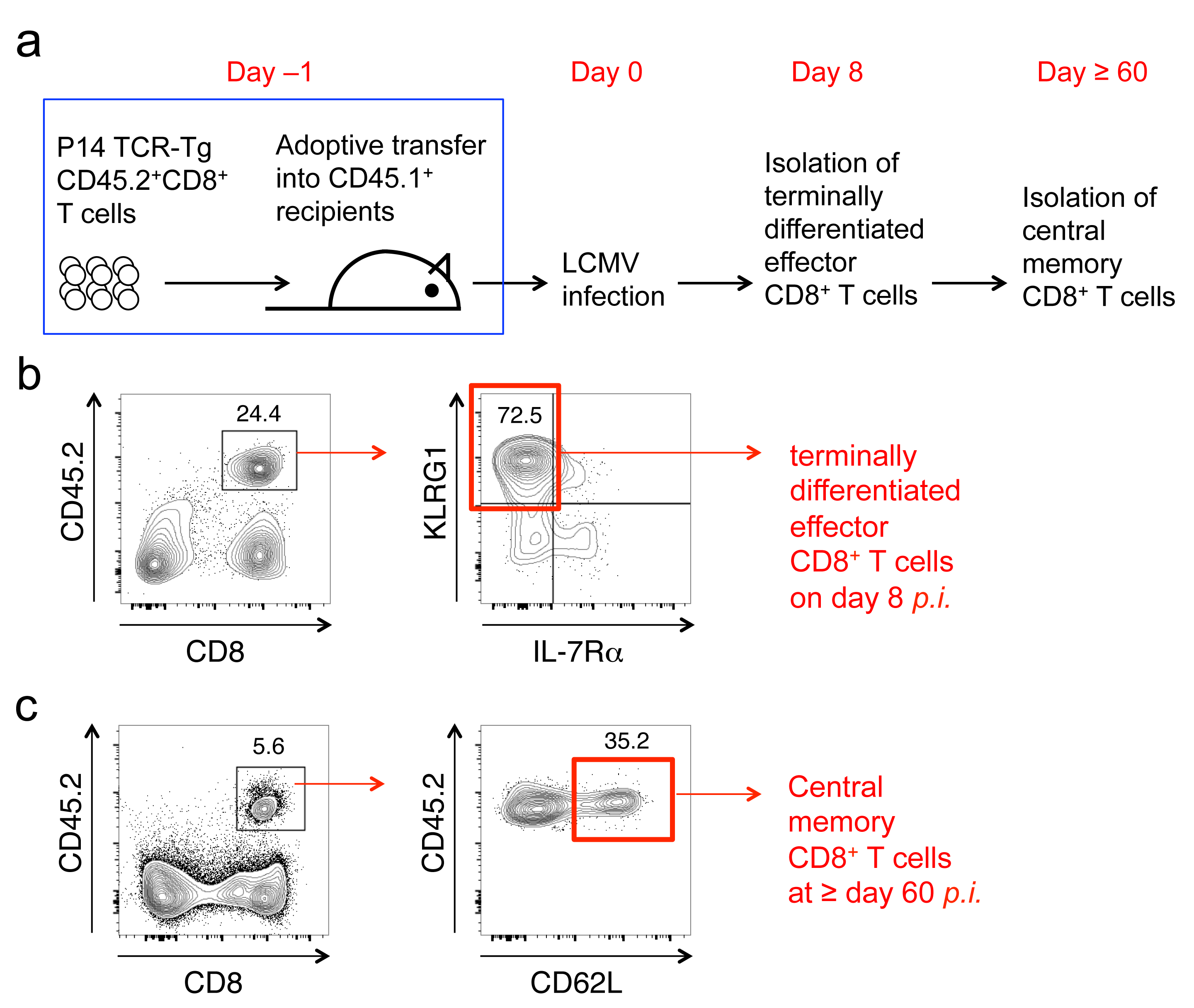
Differentiation of effector and memory CD8+ T cells is accompanied by extensive changes in the transcriptome and histone modifications at gene promoters; however, the enhancer repertoire and associated gene regulatory networks are poorly defined. Using histone mark chromatin immunoprecipitation coupled with deep sequencing, we mapped the enhancer and super-enhancer landscapes in antigen-specific naïve, differentiated effector, and central memory CD8+ T cells during LCMV infection. Epigenomics-based annotation revealed a highly dynamic repertoire of enhancers, which were inherited, de novo activated, decommissioned and re-activated during CD8+ T cell responses. We employed a computational algorithm to pair enhancers with target gene promoters. On average, each enhancer targeted three promoters and each promoter was regulated by two enhancers. By identifying enriched transcription factor motifs in enhancers, we defined transcriptional regulatory circuitries at each CD8+ T-cell response stage. These multi-dimensional datasets provide a blueprint for delineating molecular mechanisms underlying functional differentiation of CD8+ T cells. |
|
Tips for navigating the supplemental tables
Supplemental Table 1. List of differentially expressed genes (DEGs) in 12 ExprClusters
Supplemental Table 2. List of DEGs newly identified in this study compared with those identified by Best et al.
Supplemental Table 3. List of differentially expressed genes (DEGs) in 10 epigenetic clusters at promoters (ProEpi Clusters).
Supplemental Table 4. List of enhancer target genes at different stages of CD8+ T cell responses.
Supplemental Table 5. List of enhancer Epigenetic (EnhEpi) clusters associated with differentially expressed genes (DEGs) during CD8+ T cell response.
Supplemental Table 6. List of super enhancers and their target genes during CD8+ T cell responses.
Supplemental Table 7. List of transcription factors, TF target genes, and GO term analysis of the target genes during CD8+ T cell responses.
Genome browser view of ChIP-Seq, RNA-Seq and Hi-C data.
RNA-Seq and ChIP-Seq data are deposited in GEO under the accession number GSE76031. Hi-C data are deposited in GEO under the accession number GSE85560.
Reference: He B, Xing S, Chen C, Gao P, Teng L, Shan Q, Gullicksrud JA, Martin MD, Yu S, Harty JT, Badovinac VP, Tan K, and Xue HH. CD8+ T cells utilize highly dynamic enhancers repertoires and regulatory circuitry in response to infections. 2016. Immunity.
Send questions and comments to Kai Tan (tank1@email.chop.edu) or Hai-Hui Xue (hai-hui-xue@uiowa.edu).
